Automating Organ Segmentation Using Deep Learning

This is the first in a series of two posts detailing the work our fabulous summer interns did in 2021. Want to join us for the summer of 2022? Apply to join us!
This past summer two of our interns, David Wu and Chemda Wiener, developed a deep learning model to automatically segment lungs from computed tomography (CT) imagery.
Segmenting internal organs is crucial to treating cancer patients with radiotherapy. In radiotherapy, beams of radiation are used to kill cancerous cells. The better the segmentation, the more targeted the dose of radiotherapy can be, thereby minimizing damage to healthy tissue.
By automating this process we can improve healthcare clinician efficiency and ensure consistency in quality of treatment and reduce variation in performance. Segmentation performed manually
Model architecture
To develop our model, we utilized the U-Net architecture. U-Nets consist of two main parts: downsampling, which reduces the dimensionality of our input images and upsampling, for generating the segmentation mask. We developed a customized U-Net in Tensorflow based on the Keras U-Net package.
Loss function
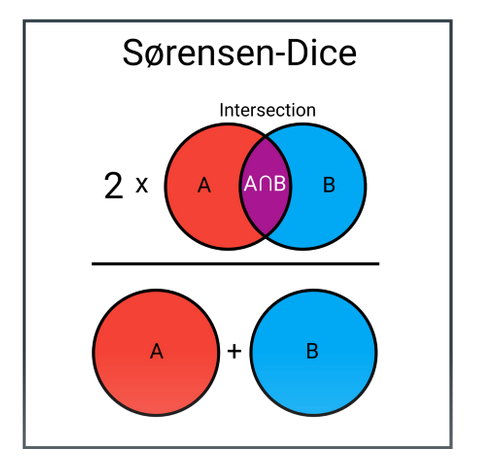
In machine learning, loss functions are critically important. They help us determine progress to our objective of optimizing performance. In our case, we utilized dice loss as our loss function. In the context of a segmentation task, dice loss measures 2 times the area of intersection between our predicted mask and the true mask, over the total area of both masks.
Resampling
CT scans come from different sites and differ in terms of voxel spacing (the physical dimensions of a pixel) and slice thickness. Resampling ensures all scans are uniform in terms of voxel spacing and thickness, aiding model performance with heterogeneous voxel spacing data. Image scans were resampled using a linear resampling transformation and masks were resampled using a nearest neighbor resampling transformation. After resampling, images were padded to provide a uniform resolution.
Evaluation metric
To evaluate the performance of our model, we used the surface-dice score. Instead of considering the overlap for all pixels in the mask, we compute loss based on the pixels that make up the edge or “surface” of the mask. We allowed a tolerance of 2mm such that differences less than that threshold are considered correct. We utilized DeepMind's surface dice implementation.
Results
Overall, using our initial model containing 60 patient series, and trained for 20 epochs we achieved a median 3d surface dice loss of 0.95 and a mean score of 0.92.

Next steps
- Intelligently crop images to optimize information density and minimize memory footprint.
- Leverage compact representations to further pursue V-Net based 3D model training.
- Expand model organ coverage to include heart, esophagus, and more.
See the Code
- See our current progress available on our open source Github repository
Special thanks
We thank Dr. Raymond Mak and Ahmed Hosny for their expertise, guidance, and tremendous support of our efforts.
Interested in working with us?
Applications are open – apply to work with us today!
